Stochastic Response¶
Richard M. Murray, 6 Feb 2022
This notebook illustrates the implementation of random processes and stochastic response. We focus on a system of the form

where 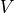 is a white noise process and the system is a first order linear system.
is a white noise process and the system is a first order linear system.
[83]:
import numpy as np
import scipy as sp
import matplotlib.pyplot as plt
import control as ct
from math import sqrt, exp
# Fix random number seed to avoid spurious figure regeneration
np.random.seed(1)
We begin by defining a simple first order system

and a (scalar) white noise signal 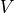 with intensity
with intensity 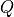 .
.
[84]:
# First order system
a = 1
c = 1
sys = ct.ss([[-a]], [[1]], [[c]], 0)
# Create the time vector that we want to use
Tf = 5
T = np.linspace(0, Tf, 1000)
dt = T[1] - T[0]
# Create the basis for a white noise signal
Q = np.array([[0.1]])
V = ct.white_noise(T, Q)
plt.plot(T, V[0])
plt.xlabel('Time [s]')
plt.ylabel('$V$');

Note that the magnitude of the signal seems to be much larger than 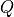 . This is because we have a Guassian process
. This is because we have a Guassian process 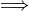 the signal consists of a sequence of “impulse-like” functions that have magnitude that increases with the time step
the signal consists of a sequence of “impulse-like” functions that have magnitude that increases with the time step 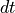 as
as 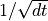 (this gives covariance
(this gives covariance 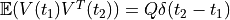 .
.
[85]:
# Calculate the sample properties and make sure they match
print("mean(V) [0.0] = ", np.mean(V))
print("cov(V) * dt [%0.3g] = " % Q, np.round(np.cov(V), decimals=3) * dt)
mean(V) [0.0] = 0.17348786109316244
cov(V) * dt [0.1] = 0.09633133133133133
The response of the system to white noise can be computed using the input_output_response function:
[86]:
# Response of the first order system
T, Y = ct.input_output_response(sys, T, V)
plt.plot(T, Y)
plt.xlabel('Time [s]')
plt.ylabel('$Y$');
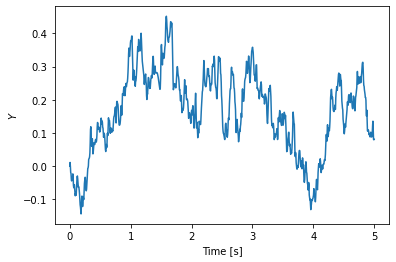
This is a first order system, and so we can compute the analytical correlation function and compare this to the sampled data:
[87]:
# Compare static properties to what we expect analytically
def r(tau):
return c**2 * Q / (2 * a) * exp(-a * abs(tau))
print("* mean(Y) [%0.3g] = %0.3g" % (0, np.mean(Y)))
print("* cov(Y) [%0.3g] = %0.3g" % (r(0), np.cov(Y)))
* mean(Y) [0] = 0.165
* cov(Y) [0.05] = 0.0151
Finally, we look at the correlation function for the input and the output:
[88]:
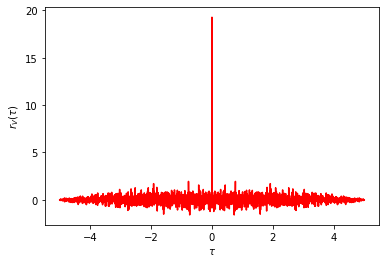
# Correlation function for the input
tau, r_V = ct.correlation(T, V)
plt.plot(tau, r_V, 'r-')
plt.xlabel(r'$\tau$')
plt.ylabel(r'$r_V(\tau)$');
[89]:
# Correlation function for the output
tau, r_Y = ct.correlation(T, Y)
plt.plot(tau, r_Y)
plt.xlabel(r'$\tau$')
plt.ylabel(r'$r_Y(\tau)$')
# Compare to the analytical answer
plt.plot(tau, [r(t)[0, 0] for t in tau], 'k--');
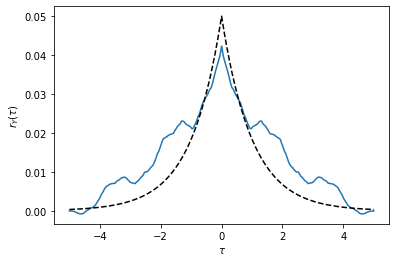
The analytical curve may or may not line up that well with the correlation function based on the sample. Try running the code again with a different random seed to see how things change based on the specific random sequence chosen at the start.
Note: the right way to compute the correlation function would be to run a lot of different samples of white noise filtered through the system dynamics and compute 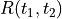 across those samples. The
across those samples. The correlation function computes the covariance between 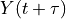 and
and  by varying
by varying 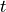 over the time range.
over the time range.
[ ]: